
- •Contents
- •Contributors
- •Part I General Principles of Cell Death
- •1 Human Caspases – Apoptosis and Inflammation Signaling Proteases
- •1.1. Apoptosis and limited proteolysis
- •1.2. Caspase evolution
- •2. ACTIVATION MECHANISMS
- •2.2. The activation platforms
- •2.4. Proteolytic maturation
- •3. CASPASE SUBSTRATES
- •4. REGULATION BY NATURAL INHIBITORS
- •REFERENCES
- •2 Inhibitor of Apoptosis Proteins
- •2. CELLULAR FUNCTIONS AND PHENOTYPES OF IAP
- •3. IN VIVO FUNCTIONS OF IAP FAMILY PROTEINS
- •4. SUBCELLULAR LOCATIONS OF IAP
- •8. IAP–IAP INTERACTIONS
- •10. ENDOGENOUS ANTAGONISTS OF IAP
- •11. IAPs AND DISEASE
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2.1. The CD95 (Fas/APO-1) system
- •2.1.1. CD95 and CD95L: discovery of the first direct apoptosis-inducing receptor-ligand system
- •2.1.2. Biochemistry of CD95 apoptosis signaling
- •2.2. The TRAIL (Apo2L) system
- •3.1. The TNF system
- •3.1.1. Biochemistry of TNF signal transduction
- •3.1.2. TNF and TNF blockers in the clinic
- •3.2. The DR3 system
- •4. THE DR6 SYSTEM
- •6. CONCLUDING REMARKS AND OUTLOOK
- •SUGGESTED READINGS
- •4 Mitochondria and Cell Death
- •1. INTRODUCTION
- •2. MITOCHONDRIAL PHYSIOLOGY
- •3. THE MITOCHONDRIAL PATHWAY OF APOPTOSIS
- •9. CONCLUSIONS
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •3. INHIBITING APOPTOSIS
- •4. INHIBITING THE INHIBITORS
- •6. THE BCL-2 FAMILY AND CANCER
- •SUGGESTED READINGS
- •6 Endoplasmic Reticulum Stress Response in Cell Death and Cell Survival
- •1. INTRODUCTION
- •2. THE ESR IN YEAST
- •3. THE ESR IN MAMMALS
- •4. THE ESR AND CELL DEATH
- •5. THE ESR IN DEVELOPMENT AND TISSUE HOMEOSTASIS
- •6. THE ESR IN HUMAN DISEASE
- •7. CONCLUSION
- •7 Autophagy – The Liaison between the Lysosomal System and Cell Death
- •1. INTRODUCTION
- •2. AUTOPHAGY
- •2.2. Physiologic functions of autophagy
- •2.3. Autophagy and human pathology
- •3. AUTOPHAGY AND CELL DEATH
- •3.1. Autophagy as anti–cell death mechanism
- •3.2. Autophagy as a cell death mechanism
- •3.3. Molecular players of the autophagy–cell death cross-talk
- •4. AUTOPHAGY, CELLULAR DEATH, AND CANCER
- •5. CONCLUDING REMARKS AND PENDING QUESTIONS
- •SUGGESTED READINGS
- •8 Cell Death in Response to Genotoxic Stress and DNA Damage
- •1. TYPES OF DNA DAMAGE AND REPAIR SYSTEMS
- •2. DNA DAMAGE RESPONSE
- •2.2. Transducers
- •2.3. Effectors
- •4. CHROMATIN MODIFICATIONS
- •5. CELL CYCLE CHECKPOINT REGULATION
- •6. WHEN REPAIR FAILS: SENESCENCE VERSUS APOPTOSIS
- •6.1. DNA damage response and the induction of apoptosis
- •6.2. p53-independent mechanisms of apoptosis
- •6.3. DNA damage response and senescence induction
- •7. DNA DAMAGE FROM OXIDATIVE STRESS
- •SUGGESTED READINGS
- •9 Ceramide and Lipid Mediators in Apoptosis
- •1. INTRODUCTION
- •3.1. Basic cell signaling often involves small molecules
- •3.2. Sphingolipids are cell-signaling molecules
- •3.2.1. Ceramide induces apoptosis
- •3.2.2. Ceramide accumulates during programmed cell death
- •3.2.3. Inhibition of ceramide production alters cell death signaling
- •4.1. Ceramide is generated through SM hydrolysis
- •4.3. aSMase can be activated independently of extracellular receptors to regulate apoptosis
- •4.4. Controversial aspects of the role of aSMase in apoptosis
- •4.5. De novo ceramide synthesis regulates programmed cell death
- •4.6. p53 and Bcl-2–like proteins are connected to de novo ceramide synthesis
- •4.7. The role and regulation of de novo synthesis in ceramide-mediated cell death is poorly understood
- •5. CONCLUDING REMARKS AND FUTURE DIRECTIONS
- •5.1. Who? (Which enzyme?)
- •5.2. What? (Which ceramide?)
- •5.3. Where? (Which compartment?)
- •5.4. When? (At what steps?)
- •5.5. How? (Through what mechanisms?)
- •5.6. What purpose?
- •6. SUMMARY
- •SUGGESTED READINGS
- •1. General Introduction
- •1.1. Cytotoxic lymphocytes and apoptosis
- •2. CYTOTOXIC GRANULES AND GRANULE EXOCYTOSIS
- •2.1. Synthesis and loading of the cytotoxic granule proteins into the secretory granules
- •2.2. The immunological synapse
- •2.3. Secretion of granule proteins
- •2.4. Uptake of proapoptotic proteins into the target cell
- •2.5. Activation of death pathways by granzymes
- •3. GRANULE-BOUND CYTOTOXIC PROTEINS
- •3.1. Perforin
- •3.2. Granulysin
- •3.3. Granzymes
- •3.3.1. GrB-mediated apoptosis
- •3.3.2. GrA-mediated cell death
- •3.3.3. Orphan granzyme-mediated cell death
- •5. CONCLUSIONS
- •REFERENCES
- •Part II Cell Death in Tissues and Organs
- •1.1. Death by trophic factor deprivation
- •1.2. Key molecules regulating neuronal apoptosis during development
- •1.2.1. Roles of caspases and Apaf-1 in neuronal cell death
- •1.2.2. Role of Bcl-2 family members in neuronal cell death
- •1.3. Signal transduction from neurotrophins and neurotrophin receptors
- •1.3.1. Signals for survival
- •1.3.2. Signals for death
- •2.1. Apoptosis in neurodegenerative diseases
- •2.1.4. Amyotrophic lateral sclerosis
- •2.2. Necrotic cell death in neurodegenerative diseases
- •2.2.1. Calpains
- •2.2.2. Cathepsins
- •3. CONCLUSIONS
- •ACKNOWLEDGMENT
- •SUGGESTED READINGS
- •ACKNOWLEDGMENT
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •5. S-NITROSYLATION OF PARKIN
- •7. POTENTIAL TREATMENT OF EXCESSIVE NMDA-INDUCED Ca2+ INFLUX AND FREE RADICAL GENERATION
- •8. FUTURE THERAPEUTICS: NITROMEMANTINES
- •9. CONCLUSIONS
- •Acknowledgments
- •SUGGESTED READINGS
- •3. MITOCHONDRIAL PERMEABILITY TRANSITION ACTIVATED BY Ca2+ AND OXIDATIVE STRESS
- •4.1. Mitochondrial apoptotic pathways
- •4.2. Bcl-2 family proteins
- •4.3. Caspase-dependent apoptosis
- •4.4. Caspase-independent apoptosis
- •4.5. Calpains in ischemic neural cell death
- •5. SUMMARY
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. HISTORICAL ANTECEDENTS
- •7.1. Activation of p21 waf1/cip1: Targeting extrinsic and intrinsic pathways to death
- •8. CONCLUSION
- •ACKNOWLEDGMENTS
- •REFERENCES
- •16 Apoptosis and Homeostasis in the Eye
- •1.1. Lens
- •1.2. Retina
- •2. ROLE OF APOPTOSIS IN DISEASES OF THE EYE
- •2.1. Glaucoma
- •2.2. Age-related macular degeneration
- •4. APOPTOSIS AND OCULAR IMMUNE PRIVILEGE
- •5. CONCLUSIONS
- •SUGGESTED READINGS
- •17 Cell Death in the Inner Ear
- •3. THE COCHLEA IS THE HEARING ORGAN
- •3.1. Ototoxic hair cell death
- •3.2. Aminoglycoside-induced hair cell death
- •3.3. Cisplatin-induced hair cell death
- •3.4. Therapeutic strategies to prevent hair cell death
- •3.5. Challenges to studies of hair cell death
- •4. SPIRAL GANGLION NEURON DEATH
- •4.1. Neurotrophic support from sensory hair cells and supporting cells
- •4.2. Afferent activity from hair cells
- •4.3. Molecular manifestations of spiral ganglion neuron death
- •4.4. Therapeutic interventions to prevent SGN death
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •18 Cell Death in the Olfactory System
- •1. Introduction
- •2. Anatomical Aspects
- •3. Life and Death in the Olfactory System
- •3.1. Olfactory epithelium
- •3.2. Olfactory bulb
- •REFERENCES
- •1. Introduction
- •3.1. Beta cell death in the development of T1D
- •3.2. Mechanisms of beta cell death in type 1 diabetes
- •3.2.1. Apoptosis signaling pathways downstream of death receptors and inflammatory cytokines
- •3.2.2. Oxidative stress
- •3.3. Mechanisms of beta cell death in type 2 diabetes
- •3.3.1. Glucolipitoxicity
- •3.3.2. Endoplasmic reticulum stress
- •5. SUMMARY
- •Acknowledgments
- •REFERENCES
- •20 Apoptosis in the Physiology and Diseases of the Respiratory Tract
- •1. APOPTOSIS IN LUNG DEVELOPMENT
- •2. APOPTOSIS IN LUNG PATHOPHYSIOLOGY
- •2.1. Apoptosis in pulmonary inflammation
- •2.2. Apoptosis in acute lung injury
- •2.3. Apoptosis in chronic obstructive pulmonary disease
- •2.4. Apoptosis in interstitial lung diseases
- •2.5. Apoptosis in pulmonary arterial hypertension
- •2.6. Apoptosis in lung cancer
- •SUGGESTED READINGS
- •21 Regulation of Cell Death in the Gastrointestinal Tract
- •1. INTRODUCTION
- •2. ESOPHAGUS
- •3. STOMACH
- •4. SMALL AND LARGE INTESTINE
- •5. LIVER
- •6. PANCREAS
- •7. SUMMARY AND CONCLUDING REMARKS
- •SUGGESTED READINGS
- •22 Apoptosis in the Kidney
- •1. NORMAL KIDNEY STRUCTURE AND FUNCTION
- •3. APOPTOSIS IN ADULT KIDNEY DISEASE
- •4. REGULATION OF APOPTOSIS IN KIDNEY CELLS
- •4.1. Survival factors
- •4.2. Lethal factors
- •4.2.1. TNF superfamily cytokines
- •4.2.2. Other cytokines
- •4.2.3. Glucose
- •4.2.4. Drugs and xenobiotics
- •4.2.5. Ischemia-reperfusion and sepsis
- •5. THERAPEUTIC APPROACHES
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. APOPTOSIS IN THE NORMAL BREAST
- •2.1. Occurrence and role of apoptosis in the developing breast
- •2.2.2. Death ligands and death receptor pathway
- •2.2.4. LIF-STAT3 proapoptotic signaling
- •2.2.5. IGF survival signaling
- •2.2.6. Regulation by adhesion
- •2.2.7. PI3K/AKT pathway: molecular hub for survival signals
- •2.2.8. Downstream regulators of apoptosis: the BCL-2 family members
- •3. APOPTOSIS IN BREAST CANCER
- •3.1. Apoptosis in breast tumorigenesis and cancer progression
- •3.2. Molecular dysregulation of apoptosis in breast cancer
- •3.2.1. Altered expression of death ligands and their receptors in breast cancer
- •3.2.2. Deregulation of prosurvival growth factors and their receptors
- •3.2.3. Alterations in cell adhesion and resistance to anoikis
- •3.2.4. Enhanced activation of the PI3K/AKT pathway in breast cancer
- •3.2.5. p53 inactivation in breast cancer
- •3.2.6. Altered expression of BCL-2 family of proteins in breast cancer
- •5. CONCLUSION
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. DETECTING CELL DEATH IN THE FEMALE GONADS
- •4. APOPTOSIS AND FEMALE REPRODUCTIVE AGING
- •6. CONCLUDING REMARKS
- •REFERENCES
- •25 Apoptotic Signaling in Male Germ Cells
- •1. INTRODUCTION
- •3.1. Murine models
- •3.2. Primate models
- •3.3. Pathways of caspase activation and apoptosis
- •3.4. Apoptotic signaling in male germ cells
- •5. P38 MITOGEN-ACTIVATED PROTEIN KINASE (MAPK) AND NITRIC OXIDE (NO)–MEDIATED INTRINSIC PATHWAY SIGNALING CONSTITUTES A CRITICAL COMPONENT OF APOPTOTIC SIGNALING IN MALE GERM CELLS AFTER HORMONE DEPRIVATION
- •11. CONCLUSIONS AND PERSPECTIVES
- •REFERENCES
- •26 Cell Death in the Cardiovascular System
- •1. INTRODUCTION
- •2. CELL DEATH IN THE VASCULATURE
- •2.1. Apoptosis in the developing blood vessels
- •2.2. Apoptosis in atherosclerosis
- •2.2.1. Vascular smooth muscle cells
- •2.2.2. Macrophages
- •2.2.3. Regulation of apoptosis in atherosclerosis
- •2.2.4. Necrosis and autophagy in atherosclerosis
- •3. CELL DEATH IN THE MYOCARDIUM
- •3.1. Cell death in myocardial infarction
- •3.1.1. Apoptosis in myocardial infarction
- •3.1.2. Necrosis in myocardial infarction
- •3.1.3. Autophagy in myocardial infarction
- •3.2. Cell death in heart failure
- •3.2.1. Apoptosis in heart failure
- •3.2.2. Necrosis in heart failure
- •3.2.3. Autophagy in heart failure
- •4. CONCLUDING REMARKS
- •ACKNOWLEDGMENTS
- •REFERENCES
- •27 Cell Death Regulation in Muscle
- •1. INTRODUCTION TO MUSCLE
- •1.1. Skeletal muscle adaptation to endurance training
- •1.2. Myonuclear domains
- •2. MITOCHONDRIALLY MEDIATED APOPTOSIS IN MUSCLE
- •2.1. Skeletal muscle apoptotic susceptibility
- •4. APOPTOSIS IN MUSCLE DURING AGING AND DISEASE
- •4.1. Aging
- •4.2. Type 2 diabetes mellitus
- •4.3. Cancer cachexia
- •4.4. Chronic heart failure
- •6. CONCLUSION
- •SUGGESTED READINGS
- •28 Cell Death in the Skin
- •1. INTRODUCTION
- •2. CELL DEATH IN SKIN HOMEOSTASIS
- •2.1. Cornification and apoptosis
- •2.2. Death receptors in the skin
- •3. CELL DEATH IN SKIN PATHOLOGY
- •3.1. Sunburn
- •3.2. Skin cancer
- •3.3. Necrolysis
- •3.4. Pemphigus
- •3.5. Eczema
- •3.6. Graft-versus-host disease
- •4. CONCLUDING REMARKS AND PERSPECTIVES
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •29 Apoptosis and Cell Survival in the Immune System
- •2.1. Survival of early hematopoietic progenitors
- •2.2. Sizing of the T-cell population
- •2.2.1. Establishing central tolerance
- •2.2.2. Peripheral tolerance
- •2.2.3. Memory T cells
- •2.3. Control of apoptosis in B-cell development
- •2.3.1. Early B-cell development
- •2.3.2. Deletion of autoreactive B cells
- •2.3.3. Survival and death of activated B cells
- •3. IMPAIRED APOPTOSIS AND LEUKEMOGENESIS
- •4. CONCLUSIONS
- •ACKNOWLEDGMENTS
- •REFERENCES
- •30 Cell Death Regulation in the Hematopoietic System
- •1. INTRODUCTION
- •2. HEMATOPOIETIC STEM CELLS
- •4. ERYTHROPOIESIS
- •5. MEGAKARYOPOIESIS
- •6. GRANULOPOIESIS
- •7. MONOPOIESIS
- •8. CONCLUSION
- •ACKNOWLEDGMENTS
- •REFERENCES
- •31 Apoptotic Cell Death in Sepsis
- •1. INTRODUCTION
- •2. HOST INFLAMMATORY RESPONSE TO SEPSIS
- •3. CLINICAL OBSERVATIONS OF CELL DEATH IN SEPSIS
- •3.1. Sepsis-induced apoptosis
- •3.2. Necrotic cell death in sepsis
- •4.1. Central role of apoptosis in sepsis mortality: immune effector cells and gut epithelium
- •4.2. Apoptotic pathways in sepsis-induced immune cell death
- •4.3. Investigations implicating the extrinsic apoptotic pathway in sepsis
- •4.4. Investigations implicating the intrinsic apoptotic pathway in sepsis
- •5. THE EFFECT OF APOPTOSIS ON THE IMMUNE SYSTEM
- •5.1. Cellular effects of an increased apoptotic burdens
- •5.2. Network effects of selective loss of immune cell types
- •5.3. Studies of immunomodulation by apoptotic cells in other fields
- •7. CONCLUSION
- •REFERENCES
- •32 Host–Pathogen Interactions
- •1. INTRODUCTION
- •2. FROM THE PATHOGEN PERSPECTIVE
- •2.1. Commensals versus pathogens
- •2.2. Pathogen strategies to infect the host
- •3. HOST DEFENSE
- •3.1. Antimicrobial peptides
- •3.2. PRRs and inflammation
- •3.2.1. TLRs
- •3.2.2. NLRs
- •3.2.3. The Nod signalosome
- •3.2.4. The inflammasome
- •3.3. Cell death
- •3.3.1. Apoptosis and pathogen clearance
- •3.3.2. Pyroptosis
- •3.2.3. Caspase-independent cell death
- •3.2.4. Autophagy and autophagic cell death
- •4. CONCLUSIONS
- •REFERENCES
- •Part III Cell Death in Nonmammalian Organisms
- •1. PHENOTYPE AND ASSAYS OF YEAST APOPTOSIS
- •2.1. Pheromone-induced cell death
- •2.1.1. Colony growth
- •2.1.2. Killer-induced cell death
- •3. EXTERNAL STIMULI THAT INDUCE APOPTOSIS IN YEAST
- •4. THE GENETICS OF YEAST APOPTOSIS
- •5. PROGRAMMED AND ALTRUISTIC AGING
- •SUGGESTED READINGS
- •34 Caenorhabditis elegans and Apoptosis
- •1. Overview
- •2. KILLING
- •3. SPECIFICATION
- •4. EXECUTION
- •4.1. DNA degradation
- •4.2. Mitochondrial elimination
- •4.3. Engulfment
- •5. SUMMARY
- •SUGGESTED READINGS
- •35 Apoptotic Cell Death in Drosophila
- •2. DROSOPHILA CASPASES AND PROXIMAL REGULATORS
- •6. CLOSING COMMENTS
- •SUGGESTED READINGS
- •36 Analysis of Cell Death in Zebrafish
- •1. INTRODUCTION
- •2. WHY USE ZEBRAFISH TO STUDY CELL DEATH?
- •2.2. Molecular techniques to rapidly assess gene function in embryos
- •2.2.1. Studies of gene function using microinjections into early embryos
- •2.2.2. In situ hybridization and immunohistochemistry
- •2.3. Forward genetic screening
- •2.4. Drug and small-molecule screening
- •2.5. Transgenesis
- •2.6. Targeted knockouts
- •3.1. Intrinsic apoptosis
- •3.2. Extrinsic apoptosis
- •3.3. Chk-1 suppressed apoptosis
- •3.4. Anoikis
- •3.5. Autophagy
- •3.6. Necrosis
- •4. DEVELOPMENTAL CELL DEATH IN ZEBRAFISH EMBRYOS
- •5. THE P53 PATHWAY
- •6. PERSPECTIVES AND FUTURE DIRECTIONS
- •SUGGESTED READING
CELL DEATH IN RESPONSE TO GENOTOXIC STRESS AND DNA DAMAGE |
79 |
|
|
p53 |
|
Mdm2 |
|
|
TNFFas |
|
|
|
|
|
|
|
Puma |
14-3-3 |
p53 |
p53 |
Noxa |
|
|||
|
|
|
Bid |
p21 |
|
|
Bax |
|
|
|
Direct targeting |
|
|
|
of mitochondria |
Cell cycle arrest |
|
Apoptosis |
|
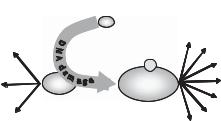
Figure 8-3. p53 is a major player influencing cell fate decisions after DNA damage. On DNA damage, p53 is stabilized and transactivates genes involved in temporary cell cycle arrest. If DNA damage persists, sustained p53 activation and/or particular post-transcriptional modifications (circle) shift p53 function toward activation of proapoptotic factors.
activity during progression through S phase, and also contributes to passage through mitosis. This leads to sustained phosphorylation and inhibition of cdk-cyclin complexes, halting cell cycle progression. Simultaneously, DNA damage activates cdk-cyclin inhibitors (p21, p27, and p57 for cdk2-cyclin E; p16 for cdk4-cyclin D or cdk6-cyclin D), leading to cell cycle arrest.
Multiple kinases activated by DNA damage (i.e., Chk2) were reported to phosphorylate and activate the tumor suppressor protein p53. In turn, p53 transcriptional activity induces proteins that contribute to both cell cycle checkpoint and DNA repair. Among those, notable is p21 transactivation (Figure 8-3), which blocks Cdc2 phosphorylation and inhibits CDK2 and CDK4, leading to G2/M and G1 arrest, respectively. p53 also transactivates 14–3-3, which sequesters Cdc25C in the cytoplasm and promotes the activation of Wee1, a tyrosine kinase that negatively regulates Cdc2, thus blocking entry into mitosis.
Additional G2/M checkpoint regulators include the mitotic serine/threonine Polo-like (Plk) and Aurora-like kinases. Plk1 promotes mitotic entry by phosphorylating and activating Cdc25C and by targeting Wee1 for degradation. In response to DNA damage, Plk1 is inhibited in an ATM/ATR-dependent manner. Inhibition of Plk1 stabilizes Wee1 and maintains Cdks in their inhibited form. Unlike Plk1, Plk3 is involved in G2/M arrest by phosphorylating and inhibiting Cdc25C.
Aurora A kinase is normally required to recruit and activate Cdk1 (cdc2)/cyclin B and to commit cells into M phase. In response to DNA damage, Aurora A kinase inhibition is associated with G2/M checkpoint activation, whereas its over-expression results in checkpoint bypass. Although early studies suggested clear, functional distinctions between DNA repair and damage checkpoint proteins, it now seems certain that many DNA damage response proteins are involved in both processes.
6. WHEN REPAIR FAILS: SENESCENCE VERSUS APOPTOSIS
Activating DNA damage checkpoints enforces growth arrest of damaged cells and allows repair mechanisms to mend the injury. Once repair is complete, cells exit the checkpoints and resume cell cycle progression and functions. However, when damage is severe or irreparable, mitotic cells from renewable tissues rely on one of two mechanisms to avoid replication. They permanently arrest the cell cycle (cellular senescence) or trigger cell death (apoptosis). These two fates are triggered by signal transduction pathways that converge on the p53 tumor suppressor (Figure 8-3). p53 activates downstream effector genes, such as those encoding the cyclin/cdk inhibitor p21 for cell cycle arrest or Puma and Bax for apoptosis. Cell fate after DNA damage depends on a balance between expression of cell cycle arrest or proapoptotic genes. The differential expression of these set of genes is regulated by two distinct, albeit not mutually exclusive, mechanisms. The first is posttranscriptional modification of p53 itself, and the second is regulation of other proteins that constrain the function of p53, such as Mdm2 or Rb. Modifications in p53 likely involve changes in p53 DNA-binding specificity or recruitment of coactivators specific for different classes of these genes. In this respect, some post-translational modifications of p53, such as phosphorylation of S46 or K120 acetylation of p53 by the histone acetyltransferases Tip60 and hMOF allows the recruitment of a coactivator required for transcription of proapoptotic genes. On the other hand, monoubiquitination or acetylation on K320 selectively favors p53 transcriptional activity on cell cycle arrest genes.
6.1. DNA damage response and the induction of apoptosis
As mentioned above, the DDR leads to phosphorylation and stabilization of p53 and subsequent upregulation of p21, which triggers G1/S arrest. For low levels of DSBs, only a minor fraction of p53 is sufficient to drive transcription of the p21 gene and cause temporary cell cycle arrest. For high levels of DSBs, p53 accumulates and in cooperation with coactivated transcription factors activates proapoptotic factors such as Bax (Bcl-2–associated X protein), PUMA (p53-upregulated modulator of apoptosis), NOXA, Bid, FAS, death receptor 5 (DR5), and PIDD (p53-induced protein with a death domain). It was also shown that the p53 protein itself targets mitochondria, where it associates with and activates the multidomain proapoptotic Bax and Bak proteins. These and others Bcl-2–like proteins exert their apoptotic effect
80 |
PABLO LOPEZ-BERGAMI AND ZE’EV RONAI |
mainly by increasing mitochondrial membrane permeabilization. Mitochondria serve as a site for convergence of multiple death-inducing stimuli, thereby serving as a pivotal decision center that controls life and death by releasing apoptogenic factors to the cytosol (intrinsic pathway). These death-inducing molecules are located within the mitochondrial intermembrane space and include cytochrome c; DIABLO/Smac, a factor that promotes caspase activation by effecting inhibitors of apoptosis proteins; the nuclease activator apoptosis-inducing factor, Endo G (an apoptotic DNase); HtrA2/Omni (an inhibitor of inhibitors of apoptosis [IAPs] that also contains proapoptotic serine protease activity); and some procaspases. In contrast, Fas and DR5 are members of the tumor necrosis factor receptor family that trigger apoptosis via the extrinsic pathway. PIDD is a caspaseactivating protein that binds adaptor protein RAIDD (RIP-associated Ich-1/CED homologous protein with death domain), which in turn binds caspase-2. Thus p53 target genes are capable of activating several apoptosis pathways, although most data argue that the mitochondrial pathway is dominant.
6.2. p53-independent mechanisms of apoptosis
Although most human tumors lose expression of functional p53, they do not lose their ability to undergo apoptosis completely. Cells employ several strategies to trigger p53-independent DNA damage-induced apoptosis. In p53 mutant tumors, the role is also played by other p53 family members such as p63 and p73. p73 is rarely mutated but often over-expressed in tumors. Whereas p53 may require p63 and p73 for triggering apoptosis, p73 is proapoptotic, even in the absence of p53. Apoptosis induced by p73 has been shown to be mediated by transcriptional upregulation of PUMA and NOXA, which in turn provokes Bax mitochondrial translocation and cytochrome c release.
Another protein that influences cellular life/death decisions independent of p53 is the mitogen-activated protein kinase (MAPK) family member c-Jun N-terminal kinase (JNK). JNK controls apoptosis both positively and negatively, depending on cell type, cell context, and stress signal. Proapoptotic JNK substrates comprise both transcription factors (e.g., c-Jun ATF-2, ATF-3, and c- Fos) and proteins that execute apoptosis (e.g., proapoptotic Bcl-2–related proteins). Recently, it was shown that JNK may also contribute to the apoptotic response of cells to activated caspases by phosphorylating H2AX at a non-canonical site that is required for apoptotic DNA fragmentation.
It has been suggested that the strength of survival signals (epidermal growth factor, insulin) determines the proor antiapoptotic effect of JNK activation. Alternatively, the role of JNK in cell fate can be regulated by a balance between cell survival signals and proapoptotic stimuli. For example, it was shown that UV-induced apoptosis is repressed by receptor tyrosine kinase–mediated inactivation of Forkhead Box O transcription factor Foxo. The function of JNK might be also governed by a timing mechanism, as shown in Drosophila. Whereas short-term activation of JNK (which is normally inhibited by a negative feedback loop involving Puc) would allow cell repair, long-term activation would lead to cell death. Such a time-dependent cellular response to JNK activation has been also observed in mammalian cells.
Whereas it is clear that JNK activation after genotoxic stress occurs through DNA damage-dependent and -independent mechanisms, the relative contribution of each of these mechanisms is still unclear. The main reason for that is that most genotoxins induce both pathways. JNK activation independent of DNA occurs when genotoxic agents activate growth factor receptors or increase reactive oxygen species levels (i.e., UV light). JNK-mediated activation by reactive oxygen species is mediated by the MAPKKK ASK1 (a specific reactive oxygen species target) and Src kinase. JNK activation by UV is also mediated by inhibition of MAPK phosphatase-1 expression.
Although JNK activation by DNA damage is well established, the mechanisms underlying the link between the DNA damage and actual activation of JNK are not as clear. Several studies have shown that JNK activation after DNA damage is mediated by ATM, DNAPKs, and Cockayne syndrome B. Alternatively, it was proposed that DNA damage per se is sufficient to elicit a signal from the nuclei to the cytosol for JNK activation. The nonreceptor tyrosine kinase c-Abl, for example, is activated after IR and is required for activation of JNK in response to cellular stress. Ras-association domain family 1C (RASSF1C) also participates in the activation of SAPK/JNK after its release to the cytoplasm from the nuclear complex containing Daxx, which is degraded by DNA damage. In all cases the signals will result in the activation of JNK or its upstream kinases MKK4/7 and MEKK1–4.
Interestingly, activating JNK, c-Jun, and ATF2 after genotoxic stress also controls the cellular response to damage by regulating the expression of various DNA repair genes, including ERCC3, XPA, RAD23B, and MSH2. Independent of its function as a transcription factor,
CELL DEATH IN RESPONSE TO GENOTOXIC STRESS AND DNA DAMAGE |
81 |
ATF2 has been revealed as an important component of the damage response. ATM phosphorylation of ATF2 has been shown to cause its localization into DNA repair foci, where it colocalizes with components of the DNA repair machinery, Rad50, NBS1, and Mre11.
Another pathway induced by DNA damage is the PI3K/Akt pathway. Activation of Akt has clearly an antiapoptotic effect that can be considered as a compensatory protective mechanism activated by the cell to escape death. Several components of the PI3K/Akt pathway have been reported to be phosphorylated in the DNA damage response, particularly Akt at Ser473. This phosphorylation, critical for Akt activity, was shown to be mediated by DNA-PK and ATM. Activated Akt then participates in an elaborate control system deciding between cell survival and death by counteracting the role of p53. Akt mediates phosphorylation and nuclear translocation of Mdm2 and inhibits interaction between Mdm2 and p19ARF, thereby potentiating the activity of Mdm2 in degrading p53. Akt can phosphorylate Chk1 and reduce the nuclear localization of Chk1, thereby interfering with Chk1-mediated p53 phosphorylation and subsequent p53 stabilization. In turn, Akt activity is controlled by p53 via transactivation of phosphatase and tensin homolog deleted in chromosome ten (PTEN) and the Akt inhibitor 14–3-3 and by repressing the catalytic subunit of PI3K.
Several evidences indicate that the transcription factor nuclear factor kappa B (NF-κB) plays a critical role in the cellular response against a variety of DNAdamaging agents. Most genotoxins transiently activate NF-κB by inducing pathways similar to those activated by cytokines (e.g., mediated by either IκB kinase complex activation or IKK/NF-κB–inducing kinase activation). On the other hand, IR and some chemotherapeutic agents induce an atypical pathway that results in a slower and more stable (approximately 4 hours) NF-κB activation that is dependent on ATM. IR activates NF- κB by increasing association of NF-κB essential modulator (NEMO) with IKKα and IKKβ, an essential step for IκB phosphorylation and subsequent degradation. The effects of DSB on NEMO are mediated by increasing NEMO sumoylation, mono-ubiquitination, and ATMdependent phosphorylation. After IκB degradation, NF- κB (mostly p50/p65 dimer) moves to the nucleus and, depending on the nature of the genotoxic signal and the cell type, regulates transcription of either proor antiapoptotic genes. UV activates another atypical pathway of activation of NF-κB. This pathway is independent of IKK and would involve Ck2-mediated phosphorylation of IκB.
In summary, DNA damage-triggered signaling and execution of apoptosis are cell-type and genotoxinspecific events that depend on p53 (p63 and p73) status, death-receptor responsiveness, MAPK activation (as well as Akt and NF-κB activation) and, most importantly, DNA repair capacity.
6.3. DNA damage response and senescence induction
Premature senescence is induced by agents that damage DNA or disrupt heterochromatin or by over-expressed oncogenes. Engagement of the DNA damage response by genotoxins was described previously in this chapter. Oncogene expression leads to elevated intracellular levels of reactive species, augmented numbers of active replicons, alterations in DNA replication fork progression, and the appearance of DNA singleand doublestrand breaks that initiate damage responses.
Senescent cells are typically characterized by a large, flat morphology, expression of a senescence-associated β-galactosidase (SA β-gal) activity of unknown function, and senescence-associated heterochromatin foci. The foci contain modifications and associated proteins characteristic of transcriptionally silent heterochromatin, such as methylated lysine 9 of histone H3 (H3K9Me), heterochromatin protein 1 (HP1), the histone H2A variant macroH2A, and the high-mobility group A (HMGA) proteins.
Mechanistically, DNA damage-induced growth arrest depends on the functional status of Rb via the p53–p21– Rb pathway or via p53-independent Rb pathways, either p16-dependent or -independent. In senescent cells, p21 and p16 play a critical role by inhibiting CDK-dependent Rb phosphorylation, leading to transcriptional repression of E2F target genes necessary for DNA synthesis and cell cycle progression. The subsequent orchestration and temporal requirements of senescence-associated markers and cell cycle regulators such as p16INK4a, p21Cip1, CDK4, ARF, p53, or PML to induce and maintain senescence are not fully understood and seem, at least to some extent, to be context, cell type, and species dependent.
Senescence, induced by the damage response present in early premalignant tumors (likely triggered by oncogene-induced DNA damage), plays a protective function by raising a barrier to tumor progression. However, it has been speculated that senescence may produce a selective pressure that eventually favors outgrowth of malignant clones containing genetic or epigenetic defects in the genome maintenance machinery.
