
- •Иерархическая классификация гликозил-гидролаз: современное состояние и перспективы развития
- •Классификация Ферментов (IUBMB)
- •Sequence Based Classification of Glycoside Hydrolases
- •Sequence Based Classification of Glycoside Hydrolases
- •Sequence Based Classification of Glycoside Hydrolases
- •Оптическая конфигурация гликозидной связи (аксиальная или экваториальная) и механизм её гидролиза (с сохранением
- •Объединение семейств гликозидаз в кланы
- •Структура и гидролиз раффинозы
- •Утилизация сахарозы бактериями: типы -фруктозидаз
- •Подсемейства семейства GH32
- •Филогенетическое древо семейства GH32
- •Подсемейства семейства GH68
- •Множественное выравнивание последовательностей белков, представляющих семейства -фруктозидазного суперсемейства
- •Family
- •Структура -фруктозидазного (фуранозидазного) суперсемейства
- •Пространственная структура белков семейства GH32
- •Семейства гликозидаз, содержащие α-галактозидазы
- •Family GH27
- •Пространственная структура белков семейства GH27 гликозидаз
- •Family GH36
- •Четыре консервативные участка в белках α-галактозидазного суперсемейства, содержащие остатки Asp
- •Филогенетическое древо α-галактозидазного суперсемейства
- •Family
- •Rigden DJ. Iterative database searches demonstrate that glycoside hydrolase families 27, 31, 36,
- •(β/α)8-barrel-type retaining α D-glycopyranosidase families
- •Родственные связи (β/α)8 гликозидаз (Nagano et al., 2001)
- •Объединение семейств гликозидаз в кланы
- •Объединение семейств гликозидаз в кланы
- •Иерархическая классификация гликозил-гидролаз
- •Персональные гранты, поддержавшие исследования в области анализа аминокислотных последовательностей

Филогенетическое древо семейства GH32
Streptococcus mutans fructanase FruA
Bacillus subtilis ORF YveB
Arthrobacter nicotinovorans fructosyltransferase
Aspergillus niger invertase
Aspergillus niger fructosyltransferase
c
Streptococcus mutans ORF FruB
Pseudomonas mucidolens endo-inulinase
Bacillus circulans fructosyltransferase
Arthrobacter sp. inulinase
Cynara scolymus sucrose:sucrose 1-fructosyltransferase
Cichorium intybus fructan:fructan 1-fructosyltransferase |
d |
Tulipa gesneriana vacuolar invertase |
|
Vigna radiata vacuolar invertase |
|
Hordeum vulgare sucrose:fructan 6-fructosyltransferase
Allium cepa fructan:fructan 6G-fructosyltransferase
Triticum aestivum extracellular invertase
Daucus carota extracellular invertase
|
Leishmania major ORF3 |
|
|
Leishmania major ORF2 |
|
|
Leishmania major ORF1 |
|
|
Escherichia coli invertase RafD |
|
0.1 |
Escherichia coli invertase CscA |
|
Zymomonas mobilis invertase |
||
|
Bacillus sp. levanase
Paenibacillus polymyxa levanase Bacillus subtilis levanase SacC
Pseudomonas mucidolens exo-inulinase Bacteroides fragilis levanase
Tritrichomonas foetus invertase Schizosaccharomyces pombe invertase
Saccharomyces cerevisiae invertase Debaryomyces occidentalis invertase
Aspergillus ficuum inulinase Penicillium purporogenum inulinase
Aspergillus foetidus
Sucrose:sucrose 1 fructosyltransferase
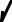
 Actinomyces naeslundii levanase
Actinomyces naeslundii levanase
 b a
b a
Thermotoga maritima inulinase/invertase
Vibrio cholerae invertase
Clostridium beijerinckii invertase
Lactobacillus plantarum invertase
Lactococcus lactis invertase Streptococcus sobrinus invertase Streptococcus mutans invertase
Staphylococcus xylosus invertase Vibrio alginolyticus invertase
Clostridium acetobutylicum invertase Geobacillus stearothermophilus invertase
Bacillus subtilis invertase Klebsiella pneumoniae invertase Salmonella typhimurium invertase

Подсемейства семейства GH68
68a |
68b |
Bacterial fructosyltransferases: |
Bacterial fructosyltransferases (levansucrases): |
Bacillus amyloliquefaciens |
Acetobacter xylinus |
Bacillus licheniformis |
Actinomyces naeslundii |
Bacillus sp. |
Arthrobacter globiformis |
Bacillus subtilis |
Arthrobacter sp. |
Clostridium acetobutylicum |
Burkholderia ambifaria |
Geobacillus stearothermophilus |
Burkholderia cenocepacia |
Lactobacillus acidophilus |
Burkholderia mallei |
Lactobacillus gasseri |
Burkholderia pseudomallei |
Lactobacillus johnsonii |
Burkholderia vietnamiensis |
Lactobacillus reuteri |
Clostridium acetobutylicum |
Lactobacillus sanfranciscensis |
Erwinia amylovora |
Leuconostoc citreum |
Erythrobacter litoralis |
Leuconostoc mesenteroides |
Gluconacetobacter diazotrophicus |
Paenibacillus polymyxa |
Gluconacetobacter xylinus |
Streptococcus mutans |
Gluconobacter oxydans |
Streptococcus salivarius |
Haloarcula marismortui |
|
Leuconostoc mesenteroides |
|
Novosphingobium aromaticivorans |
|
Pseudomonas aurantiaca |
|
Pseudomonas syringae |
|
Rahnella aquatilis |
|
Sphingomonas chungbukensis |
|
Zymomonas mobilis |
|
Extracellular invertase Zymomonas mobilis |
|
|

Множественное выравнивание последовательностей белков, представляющих семейства -фруктозидазного суперсемейства
P49943 |
84 |
LWDCDVVCK |
--DG-- |
KYYMYFP<34>I-DPAVWDD--- |
GDGNYYIYFGG<66>RFFEASWMHKYNG-------- |
KYYFSYST< 8 >ATGDN-PYGPFT |
|||
P42256 |
88 |
LWAPDVSYV-- |
DG-- |
LYYVYYA<42>I-DPNLIYA---- |
DGTYYINFGS<85>HAEEGSYMFQYGD-------- |
YYYLFYSA<21>CRSTS-PTGDFV |
|||
O52575 |
72 |
IWAPDLSYA-- |
DG-- |
KFWLIYT<36>F-DASLFHD--- |
PSGKKYLVNMY<38>AYTEGPHLYYIND-------- |
MYYLMTAE<11>ARSKT-IHGPYE |
|||
P45796 |
125 |
SWAPAVAHKKINGRDKFFLYFA<40>F-DPAVLVD--- |
DDGTGYLYSGG<39>YLFEDSGIHKYNG-------- |
KYYYSYCI<17>MVSDN-PMGPFT |
|||||
CAB66926 |
252 |
GVAISDNP--- |
TG-- |
PFDYLYS< 8 >R-DMTVYKD--- |
DDNVAYLIYSS<28>QHREAPAIFKHQN-------- |
TYYMITSG<11>HAAES-IMGPWE |
|||
AAD32559 |
603 |
IWVLAYQWG-- |
SW-- |
PFSYRTS<29>I-DQTLIAD---- |
DKNMYLFFAG<32>NLFEAPQVYKVQG----- |
QNQYFMIVEAR<10>FTATS-LNGPWT |
|||
P96463 |
263 |
IWVLGYQWG-- |
SW-- |
PFIYRTS<27>I-DQTLIAD---- |
GQNMYLFFAG<32>NLFEGVQVYKVQG----- |
QNQYLMIVEAM< 9 >FTASS-LSGSWT |
|||
BAA85252 |
116 |
VWILAYQWG-- |
PT-- |
SFSYKTS<27>I-DQTVIGD---- |
DTDMYLFFAG<32>NLFEAVQVYTVDG----- |
QNKYLMIVEAM< 9 >FTADS-LDGEWT |
|||
P79021 |
116 |
IWVLAYQWG-- |
SS-- |
TFTYRTS<27>I-DQTVIGD---- |
DTNMYLFFAG<32>DLFEAVQVYTVDG--GEGDTKYLMIVEAI<10>FTASS-LGGEWT |
||||
P23031 |
411 |
KWYLITQWA--- |
G---- |
AYATT<22>L-DFWVICN---- |
DTHCYLYFSR<34>YLFEAANVYKLDG----- |
QNRYLLMVEAY< 5 >FSAPG-QRPAWM |
|||
AAD36914 |
28 |
-------------------------- |
|
|
F-NPAVVYD---- |
GELFHLLYRA<36>YGVEDPRITKIGD------- |
EYYMVYTAY<34>LFPEK-INGKYV |
||
CAB50037 |
23 |
-------------------------- |
|
|
Y-NPAVIRE---- |
GSTYVMLYRG<37>MGVEDPRIVKIGK------- |
DYLMTYTGY<43>ILPEK-VSGEYV |
||
AAD35864 |
49 |
-------------------------- |
|
|
F-NPGAVLV---- |
GKALHVFPRL<44>LGCEDPRVFFRNS------- |
RFELLYTAK<32>ISIKS-TLGEYV |
||
BAA79294 |
61 |
-------------------------- |
|
|
F-NSTLRLA---- |
REGLEVYGRV<47>WGAEDPRYTSVNS------- |
HDVLTYTGR<39>PLATS-IKNAFI |
||
P71783 |
148 |
-------------------------- |
|
|
S-YPWVIQD---- |
GGTYRMWYGS<41>SAACRPYVVRDAG------- |
VYRMWFCAR<38>EYPCV-FDHRGQ |
||
O33833 |
73 |
VFSGSAVEK-- |
DG-- |
KMFLVYT<45>FRDPKVNRS---- |
NGEWRMVLGS<32>KEIECPDLVRIGEKD------ |
ILIYSITS< 6 >SMGEL-KEGKLN |
|||
P13394 |
109 |
VYSGGALVE-- |
NN-- |
QVLLFFT<40>FRDPKVWKK---- |
GDDYLMVVGA<34>YMWECPDFFEINGQS------ |
VMLFSPQG<12>YSVAY-IVGDQL |
|||
P05656 |
104 |
IFSGSAVVD-- |
KN-- |
NTSGFQT<48>FRDPKVFWY-- |
EKEKKWVMVLAA<29>GVWECPDLFELPVDGNPNQKKWVMQVSVG< 7 >SGMQY-FVGDFD |
||||
S33920 |
120 |
VFDGSVIPSGING-- |
LPTLLYT<52>FRDPYVFQN<9>KNNTWYTVISG<47>FNFETGNVFSLDEYGYNPHGQIFSTIGTE<13>HDMLW-VSGNVS |
||||||
P26792 |
132 |
CRSGSATILP-GN-- |
KPVILYT<48>FRDPTTAWL-- |
DKSGHWKMLVGS<32>GMWECPDFFPVSLKGLNG |
-----------<22>RYEYY-TVGTYL |
||||
P05655 |
162 |
EWSGSATFTS-DG-- |
KIRLFYT<64>LRDPHYVED---- |
KGHKYLVFEA<75>DEIERANVFKMNG------- |
KWYLFTDSR<18>YVSNS-LTGPYK |
||||
Q55242 |
354 |
QWSGSATVNS-DG-- |
SIQLYYT<74>LRDPHIIED---- |
NGSRYLIFES<78>DEVERPSVVKMGG------- |
KYYLFTASR<25>FVSDS-LRGKYR |
||||
Q43998 |
123 |
EWSGSSRLMQIHG-NTVSVFYT<63>FRDPFTFED-PKHPGVNYMVFEG<69>DQTERPQVYLHNG------- |
KYYIFTISH<16>FVGDG-IRSDFQ |
||||||
O52408 |
130 |
EWAGTPVLLNDKG-- |
DIDLYYT<51>FRDPSPFID-- |
PNDGKLYMVFEG<62>DQTERPHYVFQDG------- |
KYYLFTISH<16>FVGEH-LFGPYR |
||||
AAC36942 |
112 |
EWSGGTIMAPGSR-NQVETFFT<57>FRDPHVFIN-- |
PEDGETYALFEA<62>DQMERPHVIFQNG------- |
LTYLFTISH<16>FVSENGIFGPYE |
|||||
|
|
******** |
*********** |
********* |
********* |
***************************** |
************ |
||
GH43
GH62
GHLP
GH32
GH68
Донор H+

Family |
GH32 |
GH43 |
GH62 |
GH68 |
GHLP |
|
|
|
|
|
|
Clan |
GH-J |
GH-F |
GH-F |
GH-J |
Not known |
|
|
|
|
|
|
COG / KOG |
COG1621 / KOG0228 |
COG3507 |
None |
None |
COG2152 |
|
|
|
|
|
|
Known |
EC 2.4.1.99 |
EC 3.2.1.8 |
EC 3.2.1.55 |
EC 2.4.1.x |
Not known |
enzymatic |
EC 2.4.1.100 |
EC 3.2.1.37 |
|
EC 2.4.1.9 |
|
activities |
EC 2.4.1.x |
EC 3.2.1.55 |
|
EC 2.4.1.10 |
|
|
EC 3.2.1.7 |
EC 3.2.1.99 |
|
EC 3.2.1.26 |
|
|
EC 3.2.1.26 |
|
|
|
|
|
EC 3.2.1.65 |
|
|
|
|
|
EC 3.2.1.80 |
|
|
|
|
|
|
|
|
|
|
Molecular |
Retaining |
Inverting |
Not known |
Retaining |
Not known |
mechanism |
|
|
|
|
|
|
|
|
|
|
|
Origin |
Eukaryota: |
Eukaryota: |
Eukaryota: |
Eubacteria: |
Eukaryota: |
|
Euglenozoa |
Fungi |
Fungi |
Actinobacteria |
Fungi |
|
Fungi |
Viridiplantae |
Eubacteria: |
Firmicutes |
Metazoa |
|
Parabasalidea |
Eubacteria: |
Actinobacteria |
Proteobacteria |
Viridiplantae |
|
Viridiplantae |
Actinobacteria |
Proteobacteria |
Archaea: |
Eubacteria: |
|
Eubacteria: |
Bacteroidetes |
|
Euryarchaeota |
Actinobacteria |
|
Actinobacteria |
Firmicutes |
|
|
Aquificales |
|
Bacteroidetes |
Proteobacteria |
|
|
Cyanobacteria |
|
Chloroflexi |
Thermotogales |
|
|
Firmicutes |
|
Firmicutes |
|
|
|
Proteobacteria |
|
Fusobacteria |
|
|
|
Thermotogales |
|
Proteobacteria |
|
|
|
Archaea: |
|
Thermotogales |
|
|
|
Crenarchaeota |
|
Archaea: |
|
|
|
Euryarchaeota |
|
Euryarchaeota |
|
|
|
|
|
|
|
|
|
|

Структура -фруктозидазного (фуранозидазного) суперсемейства
|
|
|
|
|
|
|
|
GH32a |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH32b |
|
|
|
|
|
|
GH32 |
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH32c |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
clan GH-J |
|
|
|
GH32d |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH68a |
|
|
|
|
|
|
GH68 |
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH68b |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
furanosidase |
|
|
|
|
|
|
|
GH43a |
superfamily |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH43b |
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH43 |
|
GH43c |
|
|
|
|
|
|
|
|
|
|
|
|
clan GH-F |
|
|
|
GH43d |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH62 |
|
GH43e |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GH43f |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
GHLP |
|
GH43g |
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
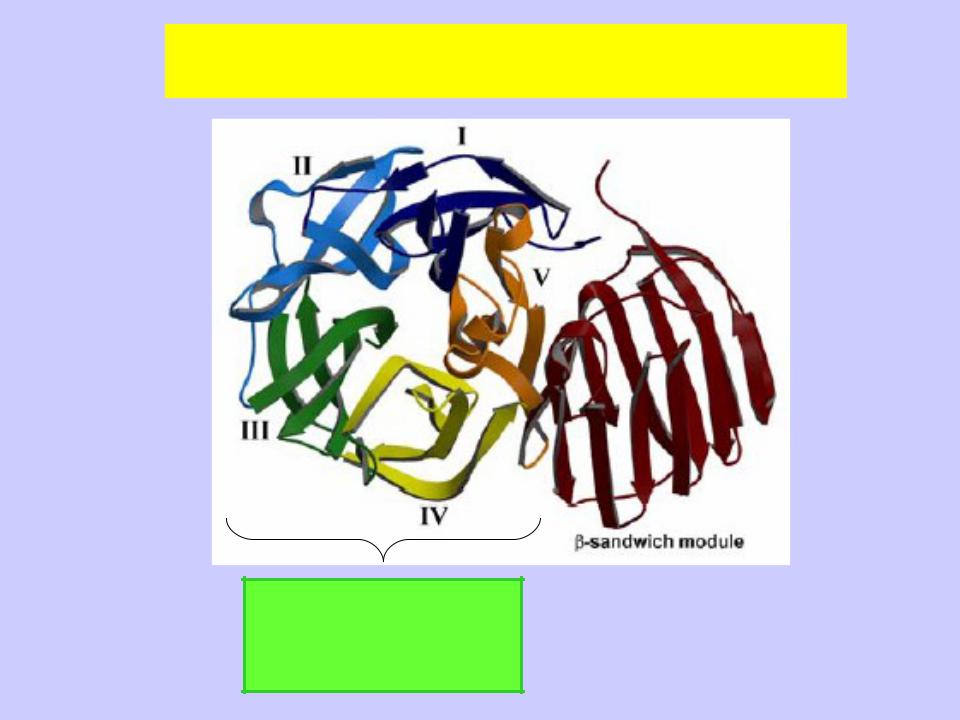
Пространственная структура белков семейства GH32
(инулиназа Thermotoga maritima)
N концевой каталитический домен в виде пяти- лопастного
β-пропеллера

EC 3.2.1.22
Common name: α-galactosidase
Reaction: Hydrolysis of terminal, non-reducing α-D-galactose residues in α D galactosides, including galactose oligosaccharides, galactomannans, and galactolipids
Molecular mechanism of hydrolyzing reaction: double displacement with overall retention of the anomeric configuration of the axial glycosidic bond
Other names: melibiase; α-D-galactosidase; α-galactosidase A; α-D-galactoside galactohydrolase; α galactoside galactohydrolase
Systematic name: α-D-galactoside galactohydrolase
Comments: Also hydrolyses α-D-fucosides

Семейства гликозидаз, содержащие α-галактозидазы
GH4 – α-galactosidase from Escherichia coli and bifunctional enzymes with α-galactosidase and α glucosidase (EC 3.2.1.20) activities from Thermotoga maritime and Thermotoga neapolitana
+6-phospho-α-glucosidases (EC 3.2.1.122) from Eubacteria
+6-phospho-β-glucosidases (EC 3.2.1.86) from Eubacteria
GH27 – mainly α-galactosidases from Eukaryota (animals, plants, fungi, etc.) and some Eubacteria
+α-N-acetylgalactosaminidases (EC 3.2.1.49) from Eukaryota (animals and fungi)
+isomalto-dextranase (EC 3.2.1.94) from Arthrobacter globiformis
GH36 – mainly α-galactosidases from Eubacteria and Eukaryota (fungi and plants)
+α-galactosyltransferases (EC 2.4.1.67 and EC 2.4.1.82) from plants
+α-N-acetylgalactosaminidase from Clostridium perfringens
+ORFs from Sulfolobus solfataricus and Sulfolobus tokodaii
GH57 – α-galactosidases from Pyrococcus furiosus and Thermococcus alcaliphilus
+ α-amylases (EC 3.2.1.1), amylopullulanase (EC 3.2.1.41), and 4-α-glucanotransferases (EC 2.4.1.x) from Eubacteria and Archaea
 clan
clan 
 GH-D
GH-D 

Family GH27
Subfamily 27a:
–plant α-galactosidases: Arabidopsis, Carica, Coffea, Cyamopsis, Glycine, Helianthus, Hordeum, Lycopersicon, Oryza, Petunia, Phaseolus, and Senna
–α-galactosyltransferase from Ajuga reptans
–animal α-galactosidases and α-N-acetylgalactosaminidases: Anopheles, Apis, Ateles, Brugia, Caenorhabditis, Ciona, Danio, Drosophila, Gallus, Homo, Mus, Rattus, Takifugu, and Tetraodon
–fungal and yeast α-galactosidases: Aspergillus, Ganoderma, Gibberella, Magnaporthe, Mortierella, Penicillium, Phanerochaete, Saccharomyces, Schizosaccharomyces, Thermomyces, Torulaspora, Trichoderma, Ustilago, and Zygosaccharomyces
–α-N-acetylgalactosaminidase from Acremonium sp.
–ORFs from Dictyostelium discoideum and Toxoplasma gondii
–bacterial α-galactosidases: Cellvibrio mixtus, Clostridium josui, Pseudomonas fluorescens, Saccharopolyspora erythraea, and Streptomyces coelicolor
–bacterial ORFs: Bacteroides, Fibrobacter, Microbulbifer, Porphyromonas, and Streptomyces
Subfamily 27b:
–plant ORFs: Arabidopsis thaliana and Oryza sativa
–bacterial ORFs: Bacillus halodurans, Bifidobacterium longum, Kineococcus radiotolerans, and Ruminococcus albus
Subfamily 27c: |
|
Unclassified proteins |
– α-galactosidase from Trichoderma reesei |
|
– isomalto-dextranase from Arthrobacter globiformis |
– ORF from Aspergillus nidulans |
|
– bacterial ORFs: Bacteroides thetaiotaomicron and Streptomyces avermitilis |
Пространственная структура белков семейства GH27 гликозидаз
(α-галактозидаза Trichoderma reesei)
GH27N
N концевой каталитический домен в виде (β/α)8-бочонка
(TIM barrel type structure)
GH27C
C концевой домен, состоящий из восьми антипараллельных β слоёв (β sandwich fold, Greek key motif)
